I am a postdoctoral fellow in the lab of Arnold Kriegstein at the University of California, San Francisco (UCSF). I focus on applying single-cell genomics techniques to study the development of specific cell types of the human brain, as well as to understand how these cell types are affected in various diseases, especially autism. Before starting my work at UCSF, I did my PhD at the University of Miami focusing on genomic analysis of autism. I did my B.S. and MS at Moscow State University in my native Russia, where I worked on animal models of epilepsy and Alzheimer’s disease.
Dmitry Velmeshev
Postdoctoral Scholar
University of California, San Francisco
From this contributor
Single-cell analysis suggests brain signaling problems in autism
Recent advances in technology allow researchers to measure RNA that is contained within the nucleus of a single brain cell.
Single-cell analysis suggests brain signaling problems in autism
Explore more from The Transmitter
‘Push-pull’ recipe for neural wiring used in multiple brain regions
A versatile pair of proteins steers neurons toward their targets and helps establish the brain’s sensory maps, new studies suggest.
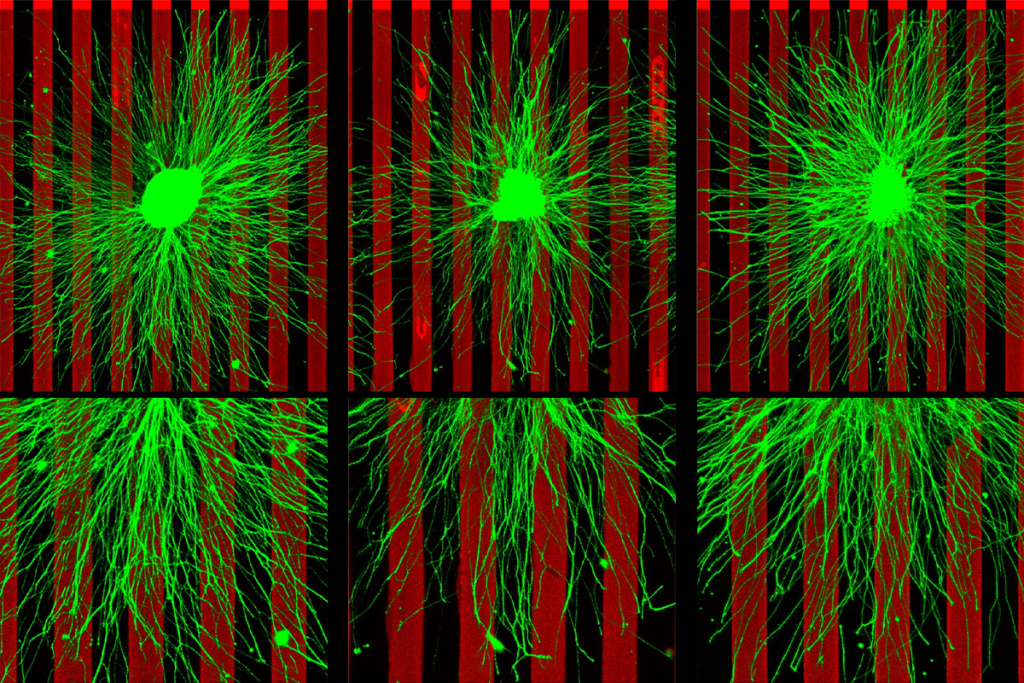
‘Push-pull’ recipe for neural wiring used in multiple brain regions
A versatile pair of proteins steers neurons toward their targets and helps establish the brain’s sensory maps, new studies suggest.
Reward-learning algorithm hardwired into dopamine circuit
The finding bolsters the canonical model of reward prediction error, which has come under scrutiny in recent years.
Reward-learning algorithm hardwired into dopamine circuit
The finding bolsters the canonical model of reward prediction error, which has come under scrutiny in recent years.
Exclusive: Brain and spinal cord institute halts research, citing funding problems
The Burke Neurological Institute, which calls itself “the only research institute in the U.S. dedicated to finding treatments to repair the brain and spinal cord,” ceased research operations on 22 May.

Exclusive: Brain and spinal cord institute halts research, citing funding problems
The Burke Neurological Institute, which calls itself “the only research institute in the U.S. dedicated to finding treatments to repair the brain and spinal cord,” ceased research operations on 22 May.